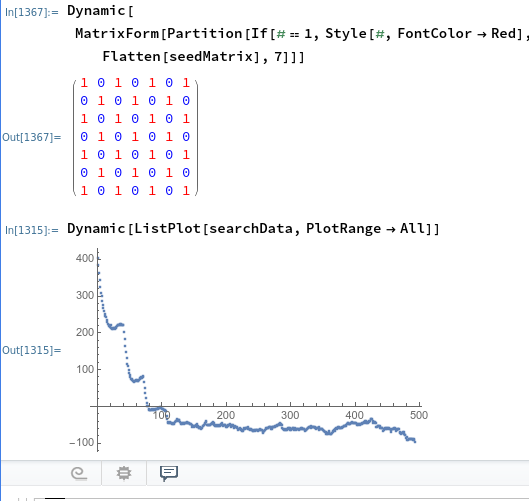
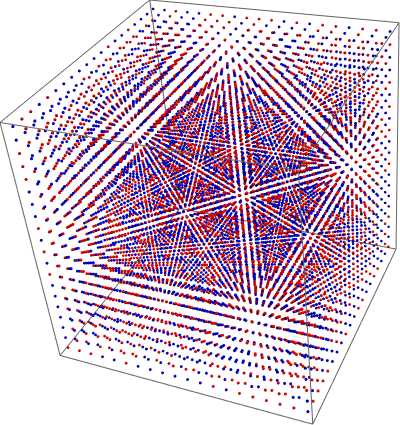
Fitting TOCSY (2D 1H NMR) Data with Mathematica

So this isn't so much "fitting" as it is programming in known data points and then making a function that easily allows one to switch between different inputs until one that is close enough is identified visually. It could be substantially improved by adding peak detection and error calculations. For one of my classes, Ch 007, my lab partner and I used solid phase synthesis techniques to make a pentapeptide incorporating four known peptides (one of which is non canonical) and an unknown mystery peptide in a known order. Solid phase techniques are generally known for their extremely high yields. Given that it was my first go at the technique I think I might have played a key role in obtaining a much lower than expected yield. Nevertheless, the product was purified by prep HPLC and the result was rather pure in the 1H NMR spectrum. We obtained a TOCSY 2D spectrum and are now tasked with interpreting the TOCSY spectrum as well as ESI-MS and MALDI-MS to verify the structur...